Home
Introduction
🧬 GENIUS-LLM
gene function inference through integrated multi-omics data with large language models
🌟 Overview
Gene function inference is critical for modern agricultural research and crop improvement. While deep learning has become the dominant computational approach, it still faces limitations in multi-omics data integration and interpretability.
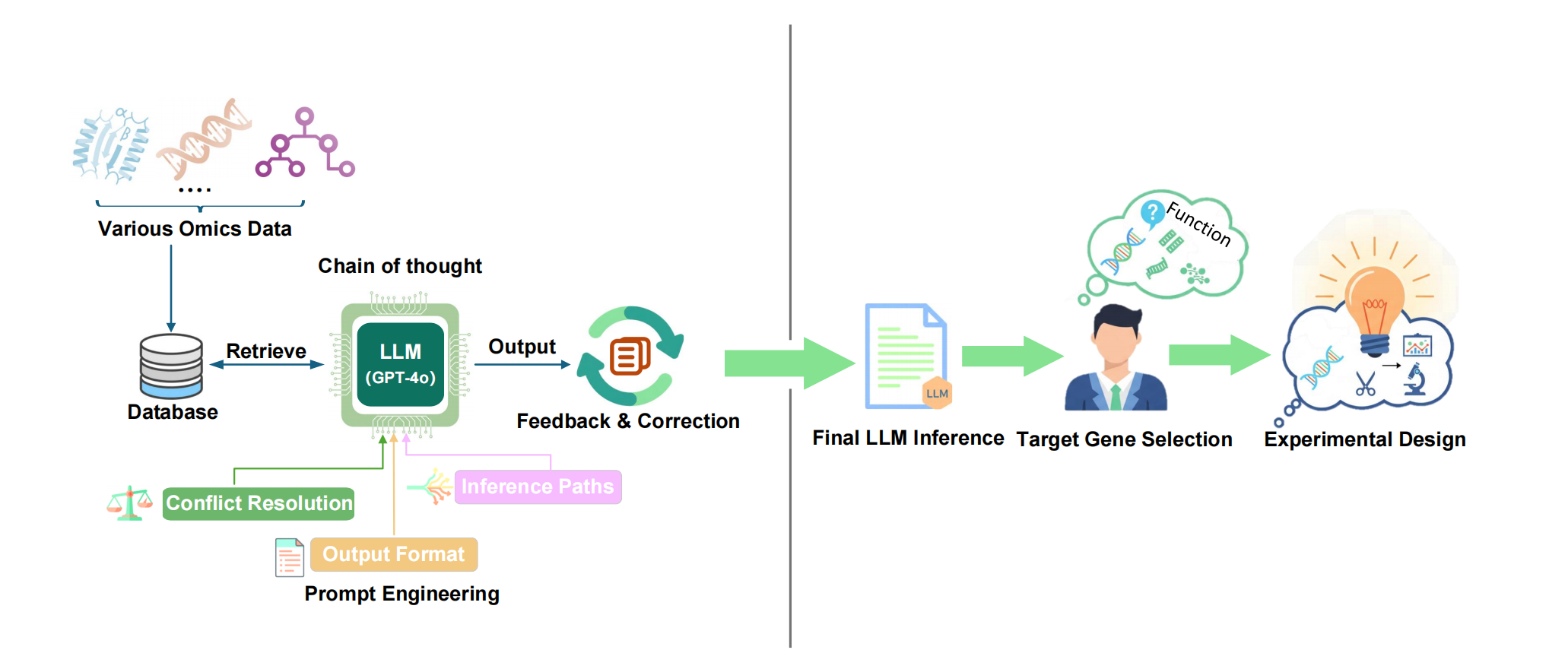
GENIUS-LLM is a one-stop platform for gene function inference that integrates multi-omics data using a Retrieval-Augmented Generation (RAG)-inspired storage system and multi-level prompt engineering. The system transforms biological data—including sequence similarity, co-expression patterns, and tissue-specific expression profiles—into structured natural language descriptions.
By utilizing priority-based analysis and Chain-of-Thought (CoT) prompting, GENIUS-LLM ensures the accuracy and interpretability of functional inference in cotton (Gossypium hirsutum), Arabidopsis thaliana, and rice (Oryza sativa).
🎯 The Nature of Output: A Hypothesis Generator
GENIUS-LLM is designed as a Hypothesis Generator:
It produces traceable, evidence-anchored, and direction-level functional hypotheses to guide downstream research.
It is not intended for novel locus discovery.
It is not designed to replace statistical mapping analyses or empirical validation.